Code
source("03.script/00_packages.R")source("03.script/00_packages.R")source("03.script/03_functions.R")Importation des deux jeux de données nécessaires à la comparaison :
(1) Les nombres de Hill issus de la diversité génétique, calculés à partir des génotypes multilocus (Hill_gen_transect)
(2) Les nombres de Hill issus de la diversité spécifique, calculés à partir des relevés floristiques (Hill_gen_transect).
Ces deux tableaux sont déjà harmonisés au niveau du transect, ce qui permet de comparer directement les deux dimensions de la diversité.
Diversité génétique
Hill_gen_transect <- read_excel(
"D:/Master 2/S10/stagem2_lcodoner/04.results/Hill_gen_transect.xlsx"
)
AllDiv <- read_excel(
"D:/Master 2/S10/stagem2_lcodoner/04.results/AllDiv.xlsx"
)
AllDiv_all <- read_excel(
"D:/Master 2/S10/stagem2_lcodoner/04.results/AllDiv_all.xlsx"
)Diversité spécifique
Hill_spe_transect <- read_excel(
"D:/Master 2/S10/stagem2_lcodoner/04.results/Hill_spe_transect.xlsx"
)Les deux jeux de données sont fusionés sur la colonne commune transect, ce qui permet d’obtenir un tableau unique contenant, pour chaque transect, les valeurs de diversité spécifique (q0_sp, q1_sp, q2_sp) et de diversité génétique (q0_gen, q1_gen, q2_gen). Cette étape gatenti que la comparaison se fait uniquement sur les transects pour lesquels les edux types de diversité sont disponibles, soit 24 transects (dits pop dans le jeu de données de génétique) sur 94. Cela évite tout biais lié à des transects manquants.
Hill_numbers <- Hill_spe_transect %>%
inner_join(Hill_gen_transect, by = "transect")– Les transects riches en espèces sont‑ils aussi riches en diversité génétique ?
– Les habitats plus stables (fixed dunes) conservent‑ils mieux la diversité génétique ?
– Observe‑t‑on un gradient nord–sud commun aux deux types de diversité ?
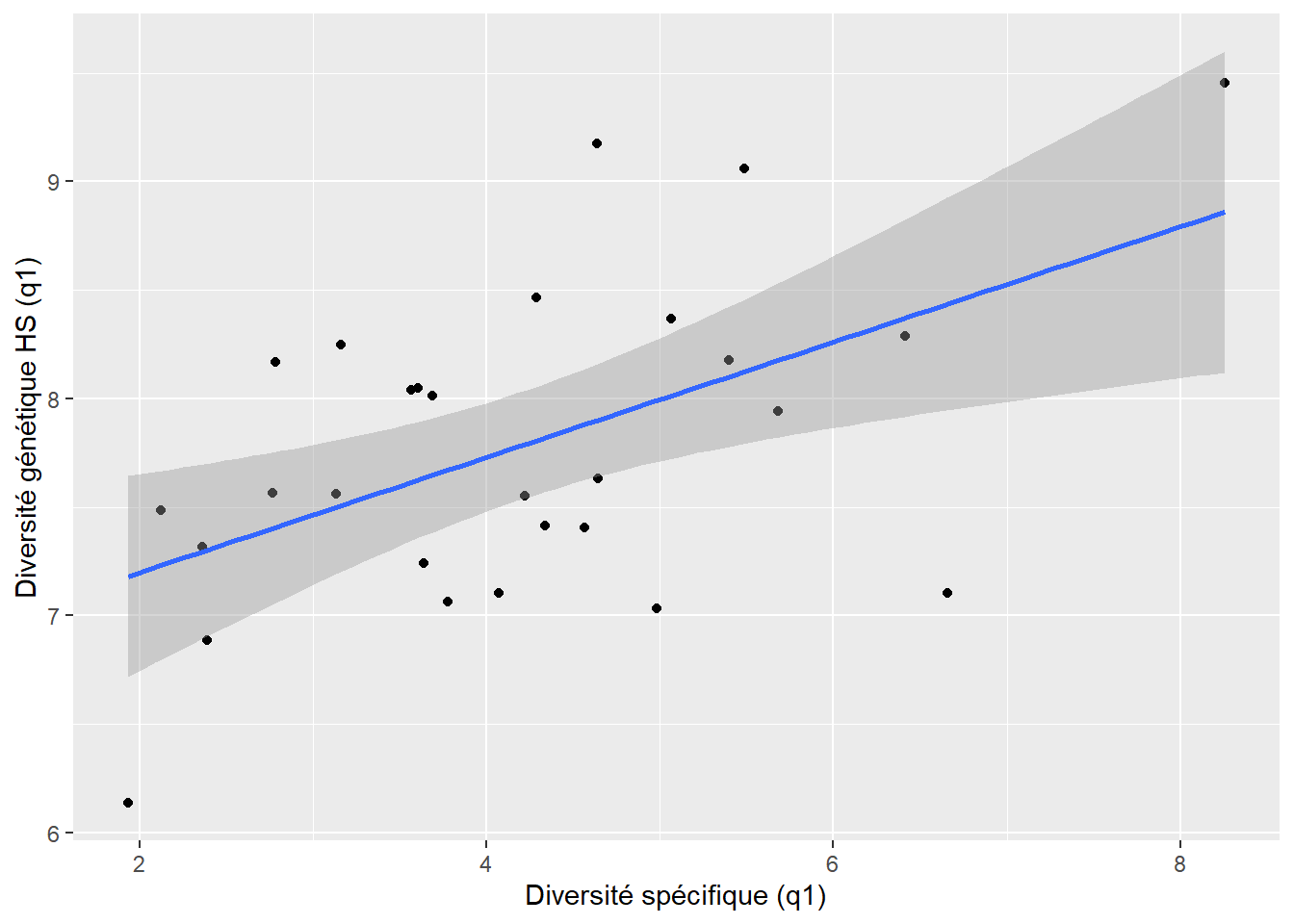
-> Ici, la diversité génétique comprend toutes les espèces confondues pour un transect. Les corrélations de Pearson sont utilisées pour évaluer la relation linéaire entre diversité spécifique et diversité génétique pour les trois ordres de Hill (q0, q1, q2). Cette analyse permet de tester si les transects riches en espèces sont également riches en diversité génétique, ou si les deux dimentsions de la biodiversité varient indépendamment.
corrélation q0
cor.test(Hill_numbers$q0_sp, Hill_numbers$q0_gen)
Pearson's product-moment correlation
data: Hill_numbers$q0_sp and Hill_numbers$q0_gen
t = -1.3954, df = 27, p-value = 0.1743
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
-0.5715345 0.1184043
sample estimates:
cor
-0.2593564 corrélation q1
cor.test(Hill_numbers$q1_sp, Hill_numbers$q1_gen)
Pearson's product-moment correlation
data: Hill_numbers$q1_sp and Hill_numbers$q1_gen
t = -2.272, df = 27, p-value = 0.03127
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
-0.66891126 -0.03998769
sample estimates:
cor
-0.400622 corrélation q2
cor.test(Hill_numbers$q2_sp, Hill_numbers$q2_gen)
Pearson's product-moment correlation
data: Hill_numbers$q2_sp and Hill_numbers$q2_gen
t = -2.502, df = 27, p-value = 0.01871
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
-0.69054458 -0.08006312
sample estimates:
cor
-0.4338384 cor_graph_q1 <- ggplot(Hill_numbers, aes(q1_sp, q1_gen)) +
geom_point(size = 3) +
geom_smooth(method = "lm", se = TRUE) +
labs(x = "Diversité spécifique (q1)", y = "Diversité génétique (q1)")`geom_smooth()` using formula = 'y ~ x'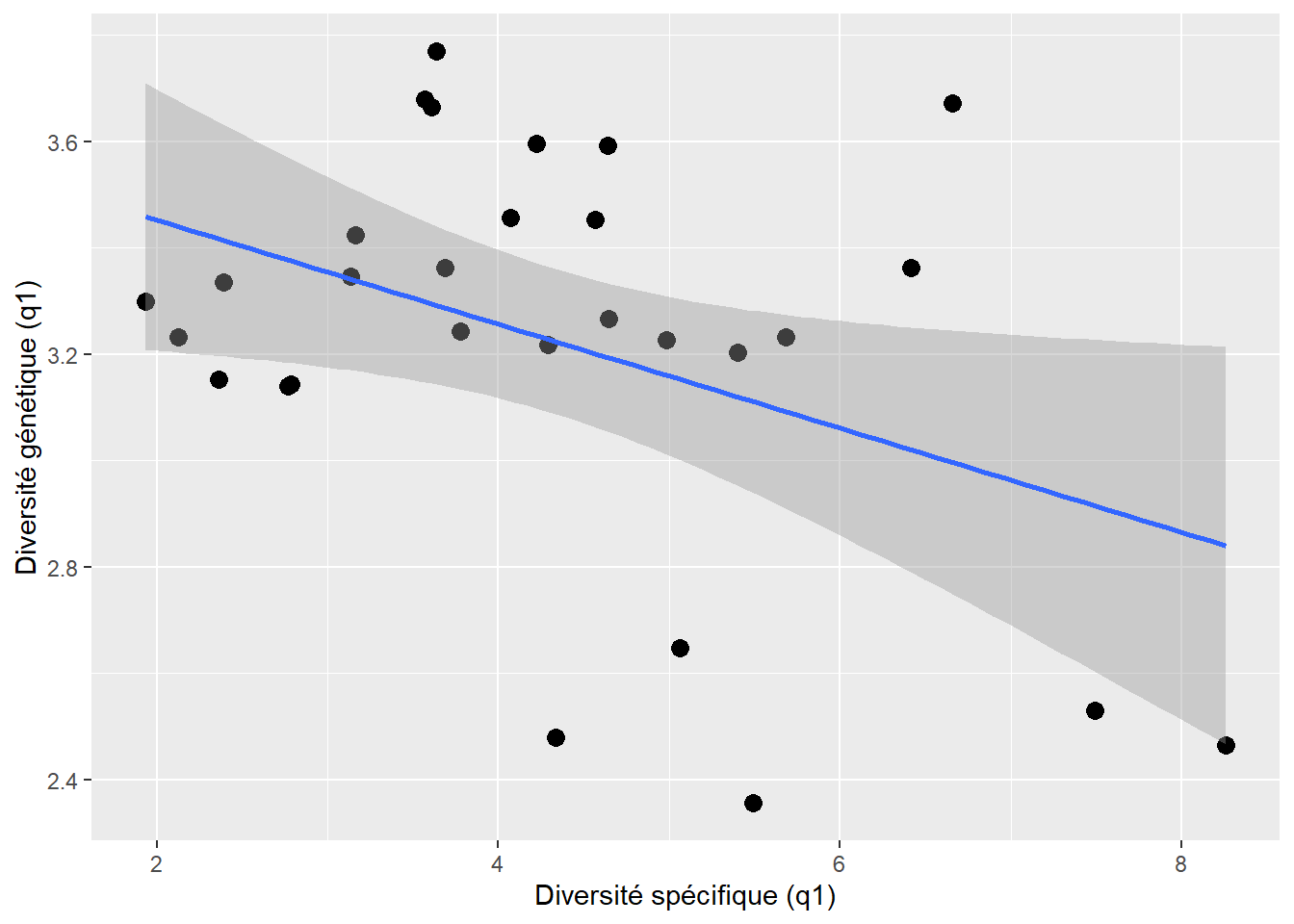
Les résultats montrent une absence de relation pour q0 (richesse), mais des corrélations négatives significatives pour q1 et q2
Des modèles linéaires simples sont ajusts pour quantifier l’effet de la diversité spécifique sur la diversité génétique.
lm(q1_gen ~ q1_sp, data = Hill_numbers)
Call:
lm(formula = q1_gen ~ q1_sp, data = Hill_numbers)
Coefficients:
(Intercept) q1_sp
3.64780 -0.09785 lm(q2_gen ~ q2_sp, data = Hill_numbers)
Call:
lm(formula = q2_gen ~ q2_sp, data = Hill_numbers)
Coefficients:
(Intercept) q2_sp
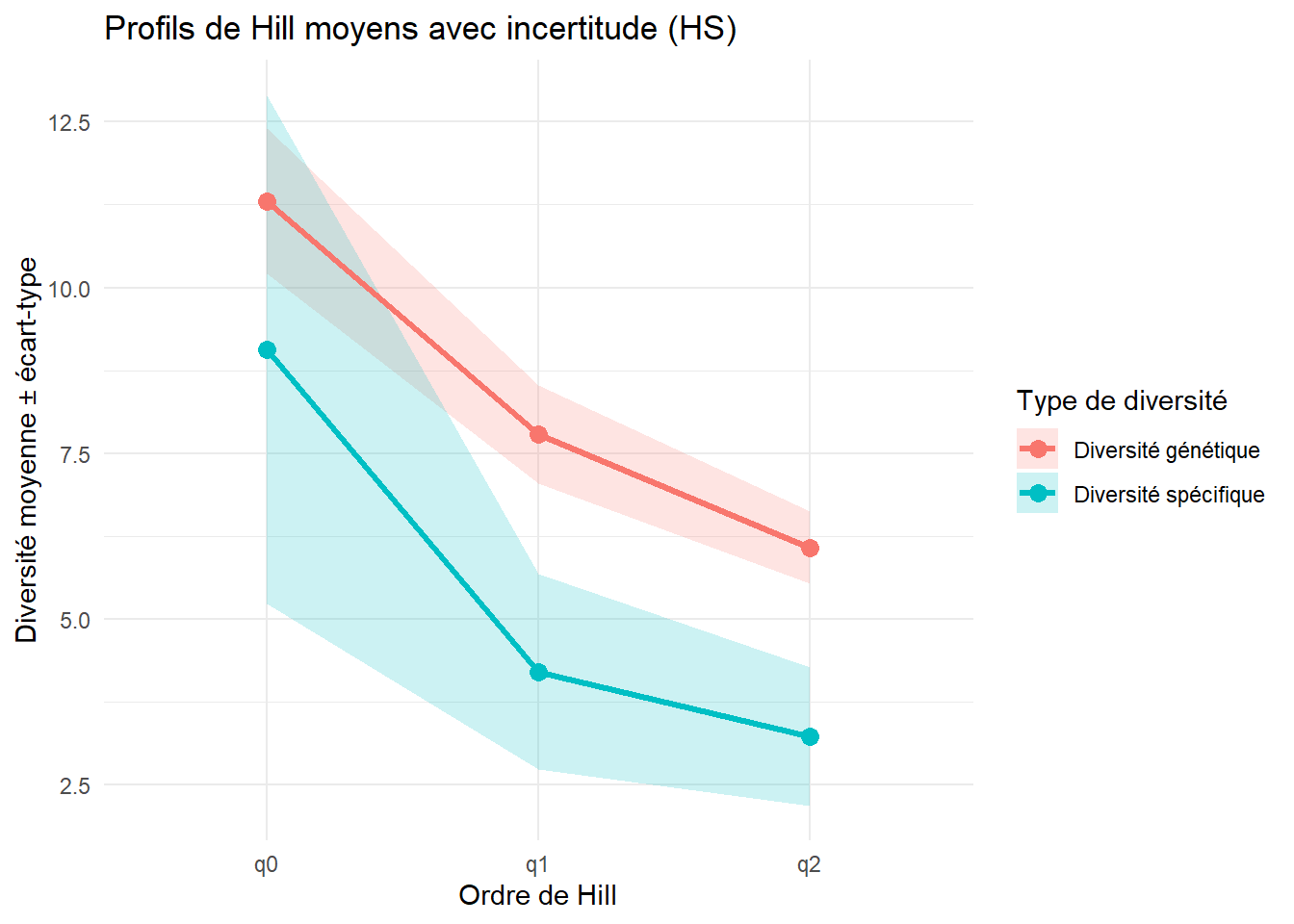
3.1278 -0.1214 Les profils de Hill sont construits pour comparer la forme des courbes de diversité spécifique et génétique. En représentant q0, q1 et q2 pour chaque transect, ces profils permettent de visualiser la structure interne de la diversité.
profils <- Hill_numbers %>%
select(transect, q0_sp, q1_sp, q2_sp, q0_gen, q1_gen, q2_gen) %>%
pivot_longer(-transect, names_to = "type", values_to = "val")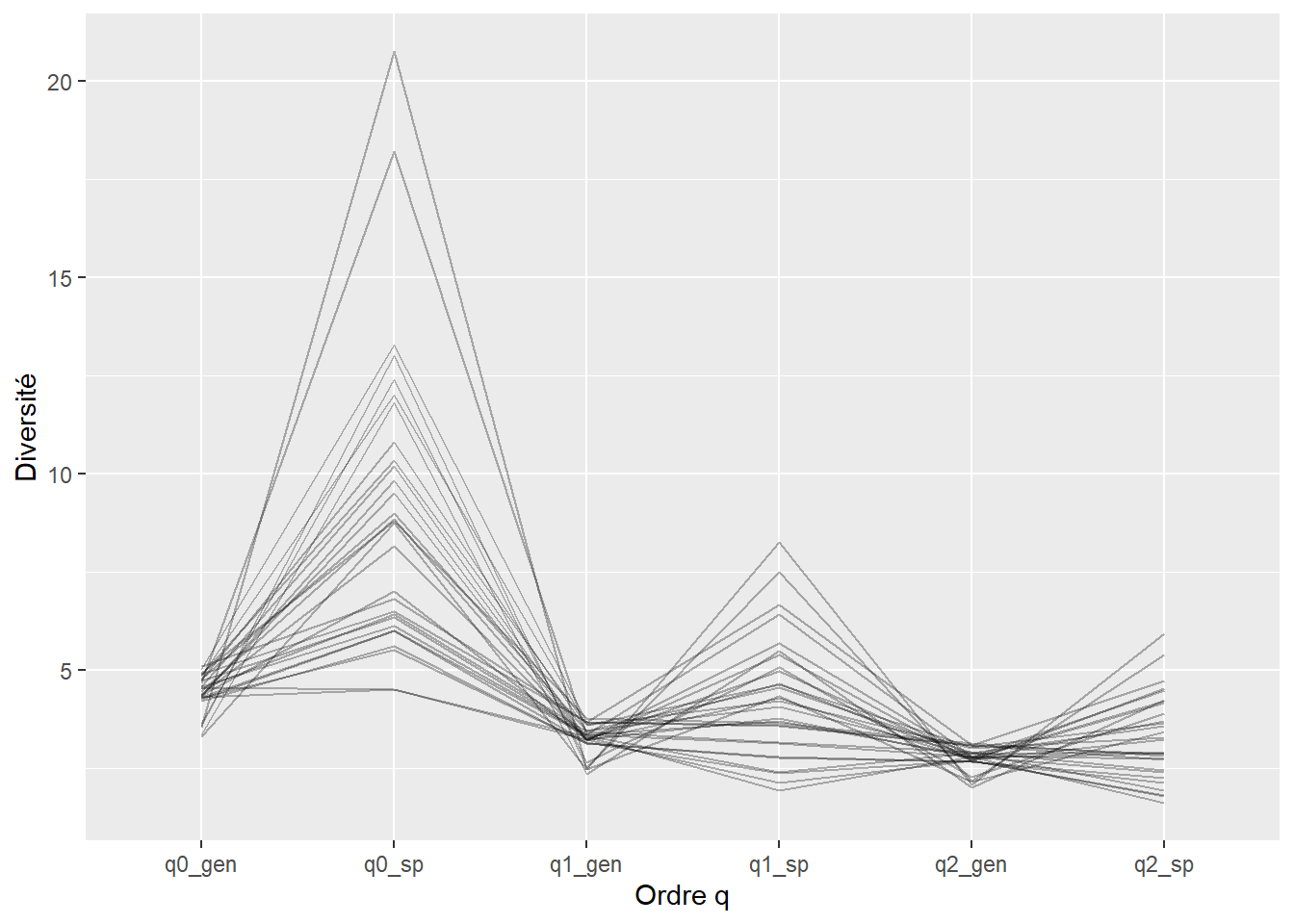
L’idée ici est de faire correspondre le tableau de diversité spécifique par transect avec la diversité génétique d’une seule espèce.
Préparation tableau diversité génétique
# Filtrer que l'espèce CV
Hill_gen_especeCS <- AllDiv %>%
filter(spe == "CS")
# Format large
Hill_gen_especeCS <- Hill_gen_especeCS %>%
tidyr::pivot_wider(
names_from = div,
values_from = val
) %>%
rename(
transect = pop,
q0_gen = q0,
q1_gen = q1,
q2_gen = q2
) %>%
select(-spe)Fusion des deux tableaux
Hill_gen_vs_speCS <- Hill_spe_transect %>%
inner_join(Hill_gen_especeCS, by = "transect")liste_especes <- unique(AllDiv_all$spe)
for (esp in liste_especes) {
cat("#### ", esp, "\n\n")
res <- analyse_espece(esp)
Hill_gen_vs_spe <- res$fusion
Hill_profils_stats <- res$profils
# --- STOCKAGE DES RÉSULTATS DE PEARSON ---
assign(
paste0(esp, "_pearson_res"),
list(
q1 = cor.test(Hill_gen_vs_spe$q1_sp, Hill_gen_vs_spe$q1_gen),
q2 = cor.test(Hill_gen_vs_spe$q2_sp, Hill_gen_vs_spe$q2_gen)
)
)
# --- STOCKAGE DU GRAPHIQUE DE CORRÉLATION ---
assign(
paste0(esp, "_cor_graph"),
ggplot(Hill_gen_vs_spe, aes(q1_sp, q1_gen)) +
geom_point() +
geom_smooth(method = "lm") +
labs(
x = "Diversité spécifique (q1)",
y = paste0("Diversité génétique ", esp, " (q1)")
)
)
# --- STOCKAGE DU PROFIL DE HILL ---
assign(
paste0(esp, "_hill_profile_stats"),
Hill_profils_stats
)
assign(
paste0(esp, "_hill_profile_graph"),
ggplot(
Hill_profils_stats,
aes(x = ordre, y = mean, color = type, group = type)
) +
geom_ribbon(
aes(ymin = mean - sd, ymax = mean + sd, fill = type),
alpha = 0.2,
color = NA
) +
geom_line(size = 1.2) +
geom_point(size = 3) +
labs(
x = "Ordre de Hill",
y = "Diversité moyenne ± écart-type",
color = "Type de diversité",
fill = "Type de diversité",
title = paste0("Profils de Hill moyens avec incertitude (", esp, ")")
) +
theme_minimal()
)
# --- EXPORT TABLEAU ---
write_xlsx(
Hill_gen_vs_spe,
path = file.path(path_res, paste0("Hill_gen_vs_spe_", esp, ".xlsx"))
)
# --- AFFICHAGE TABLEAU ---
print(datatable(Hill_gen_vs_spe))
cat("\n### Corrélations\n\n")
print(get(paste0(esp, "_pearson_res"))$q1)
print(get(paste0(esp, "_pearson_res"))$q2)
cat("\n### Graphique q1_sp vs q1_gen\n\n")
print(get(paste0(esp, "_cor_graph")))
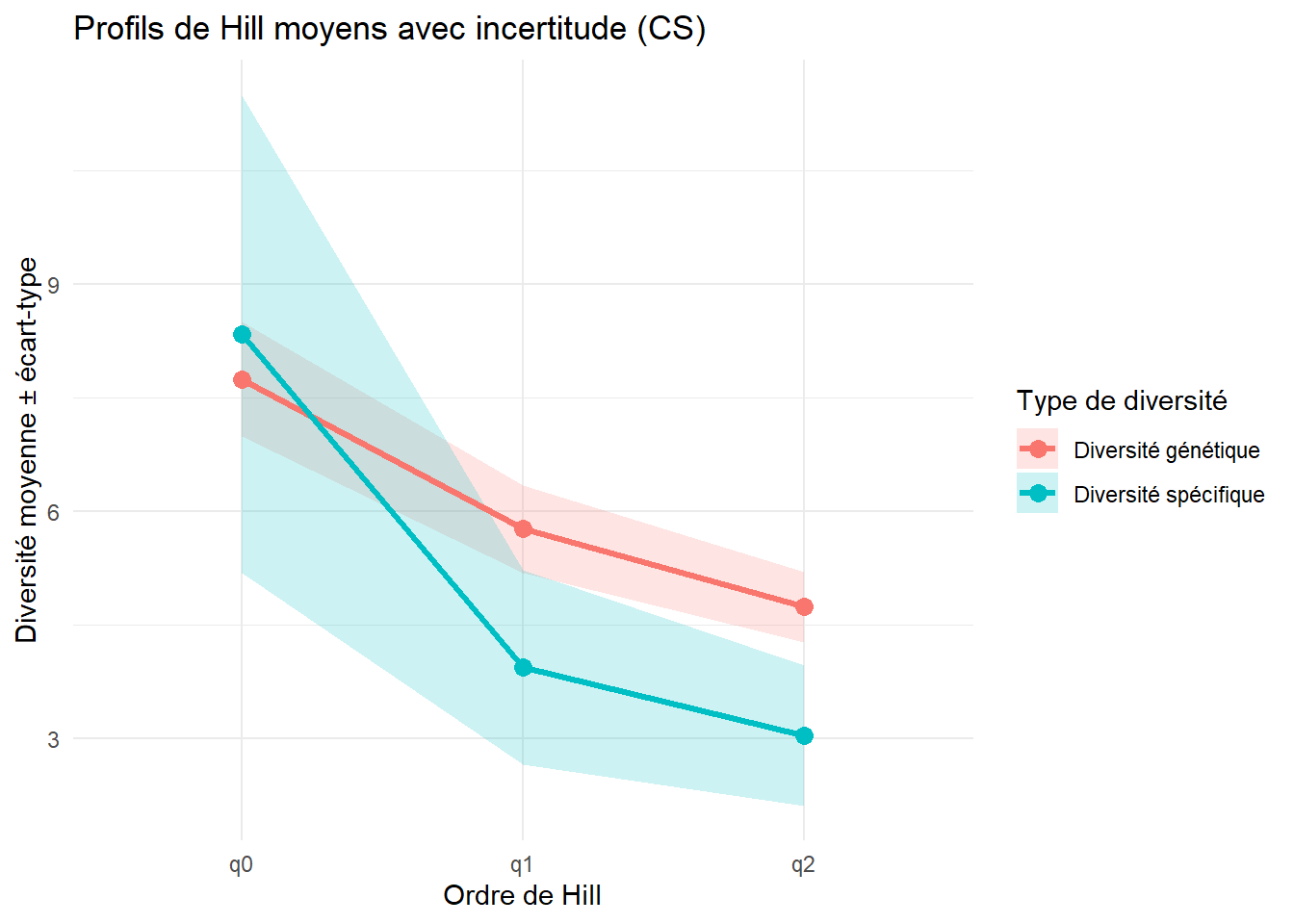
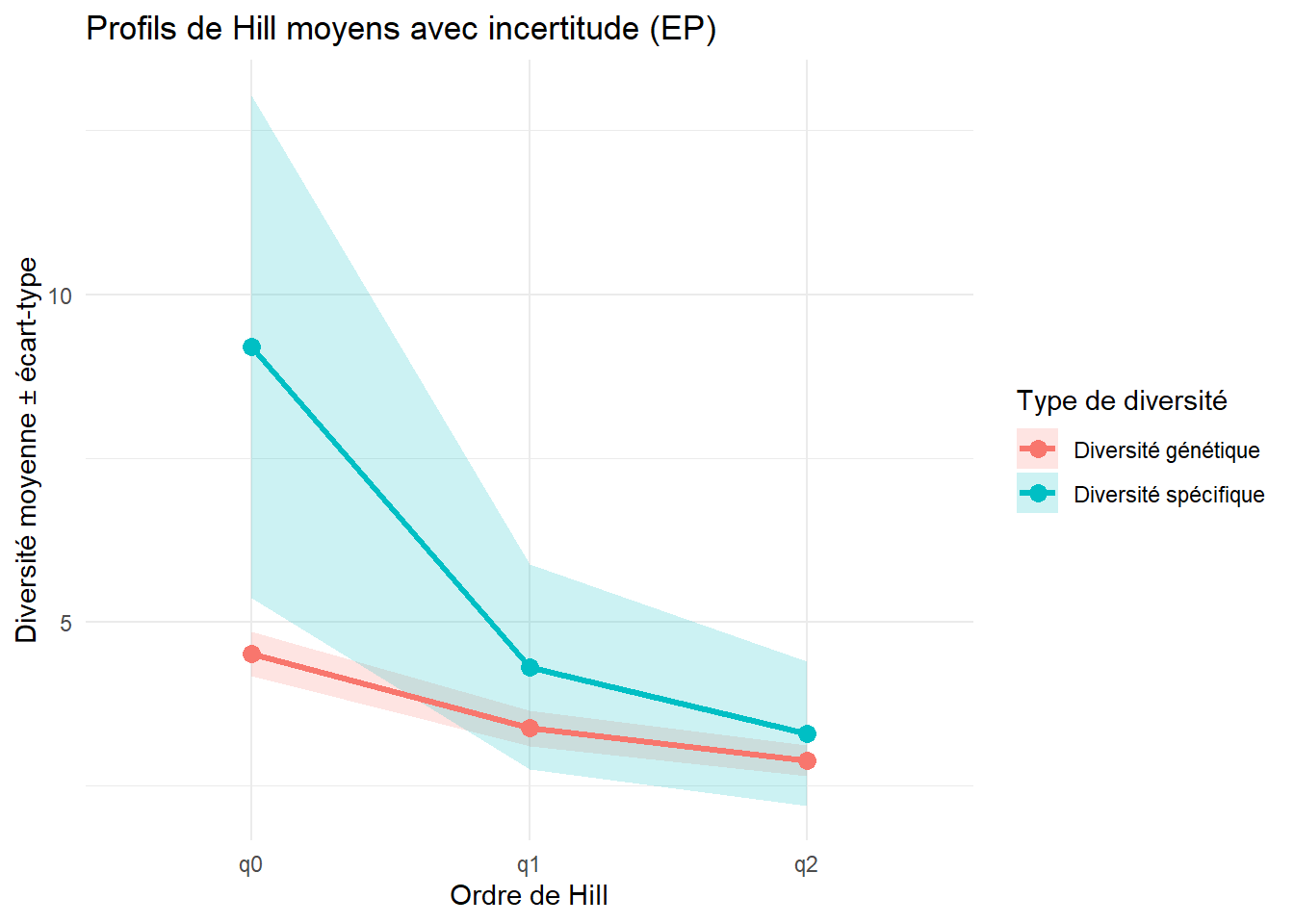
cat("\n### Profils de Hill\n\n")
print(get(paste0(esp, "_hill_profile_graph")))
cat("\n\n---\n\n")
}$q1
Pearson's product-moment correlation
data: Hill_gen_vs_spe$q1_sp and Hill_gen_vs_spe$q1_gen
t = -2.66, df = 27, p-value = 0.01299
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
-0.7045224 -0.1070508
sample estimates:
cor
-0.4556781
$q2
Pearson's product-moment correlation
data: Hill_gen_vs_spe$q2_sp and Hill_gen_vs_spe$q2_gen
t = -1.9908, df = 27, p-value = 0.05671
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
-0.64031619 0.01005131
sample estimates:
cor
-0.3577724 `geom_smooth()` using formula = 'y ~ x'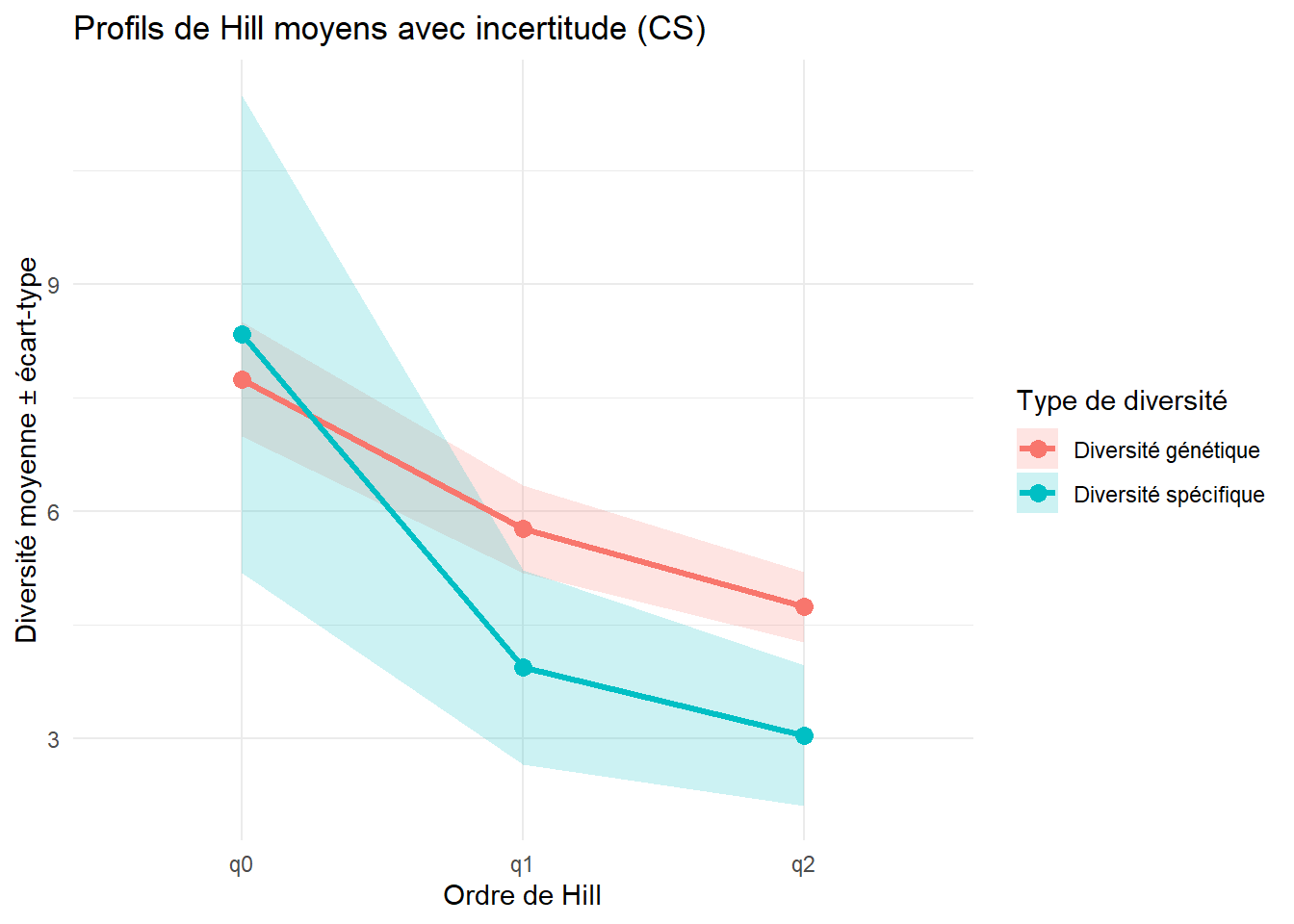
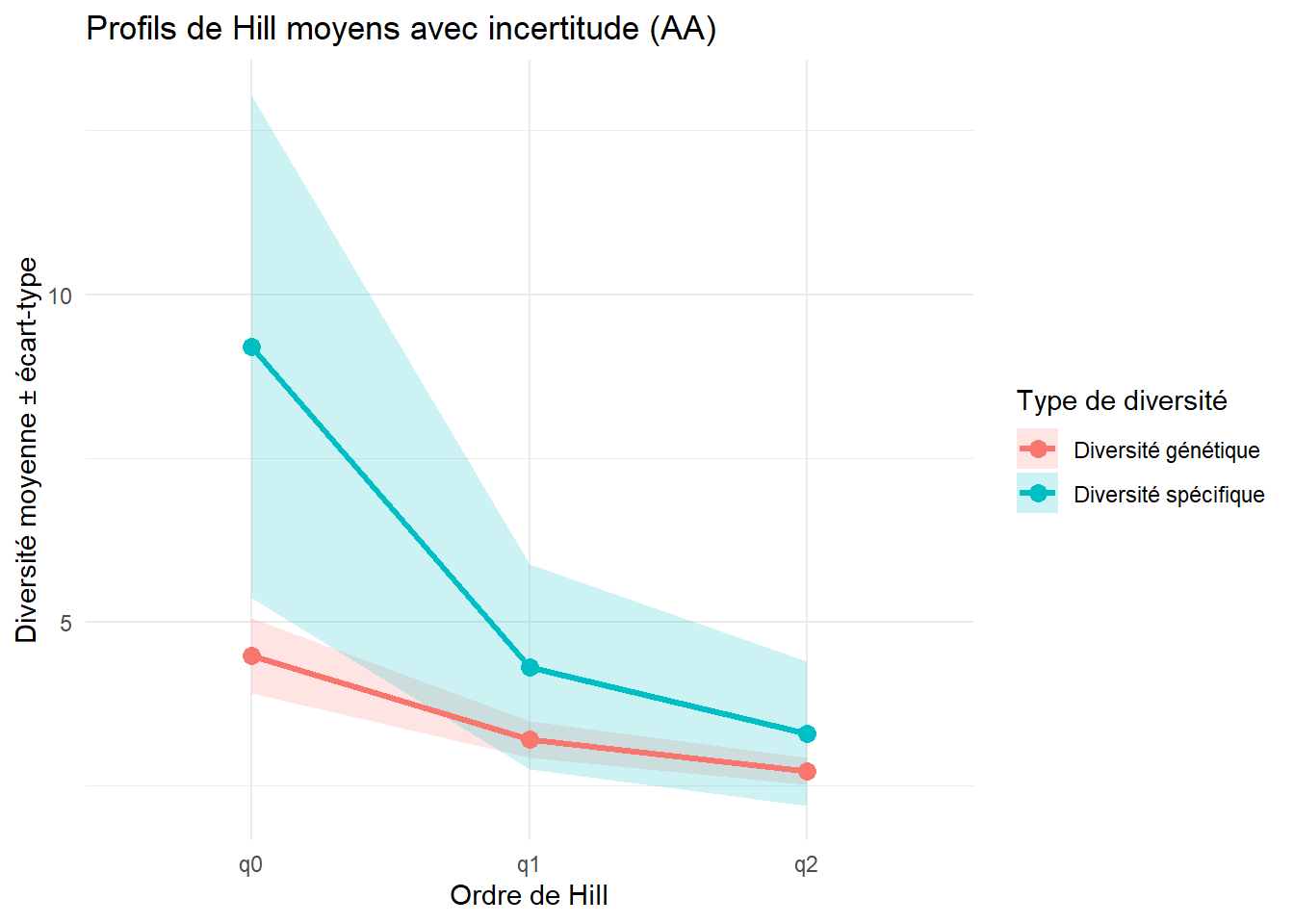
$q1
Pearson's product-moment correlation
data: Hill_gen_vs_spe$q1_sp and Hill_gen_vs_spe$q1_gen
t = 0.32976, df = 16, p-value = 0.7459
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
-0.4000551 0.5287486
sample estimates:
cor
0.08216033
$q2
Pearson's product-moment correlation
data: Hill_gen_vs_spe$q2_sp and Hill_gen_vs_spe$q2_gen
t = -0.64186, df = 16, p-value = 0.5301
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
-0.5822400 0.3330689
sample estimates:
cor
-0.1584383 `geom_smooth()` using formula = 'y ~ x'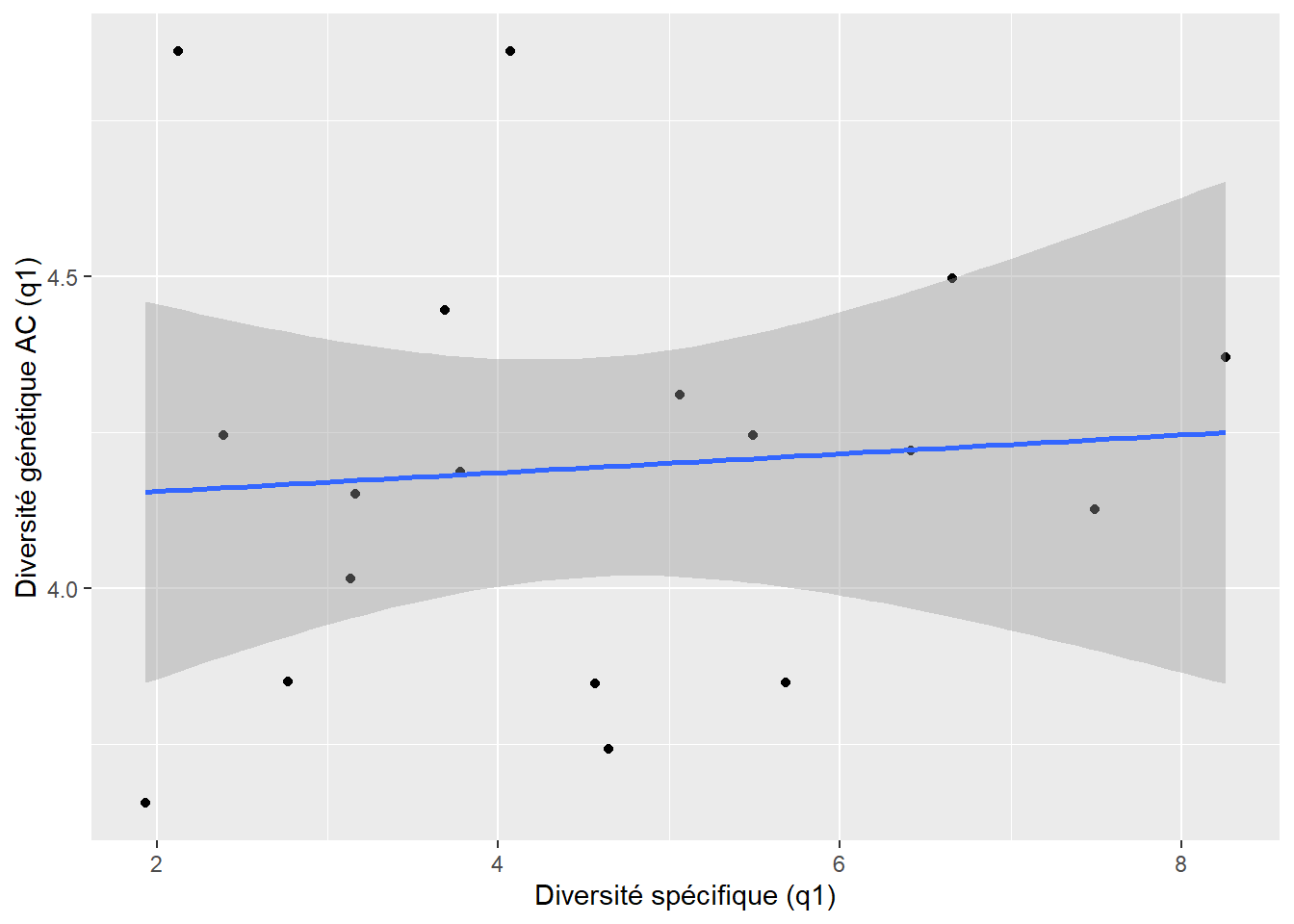
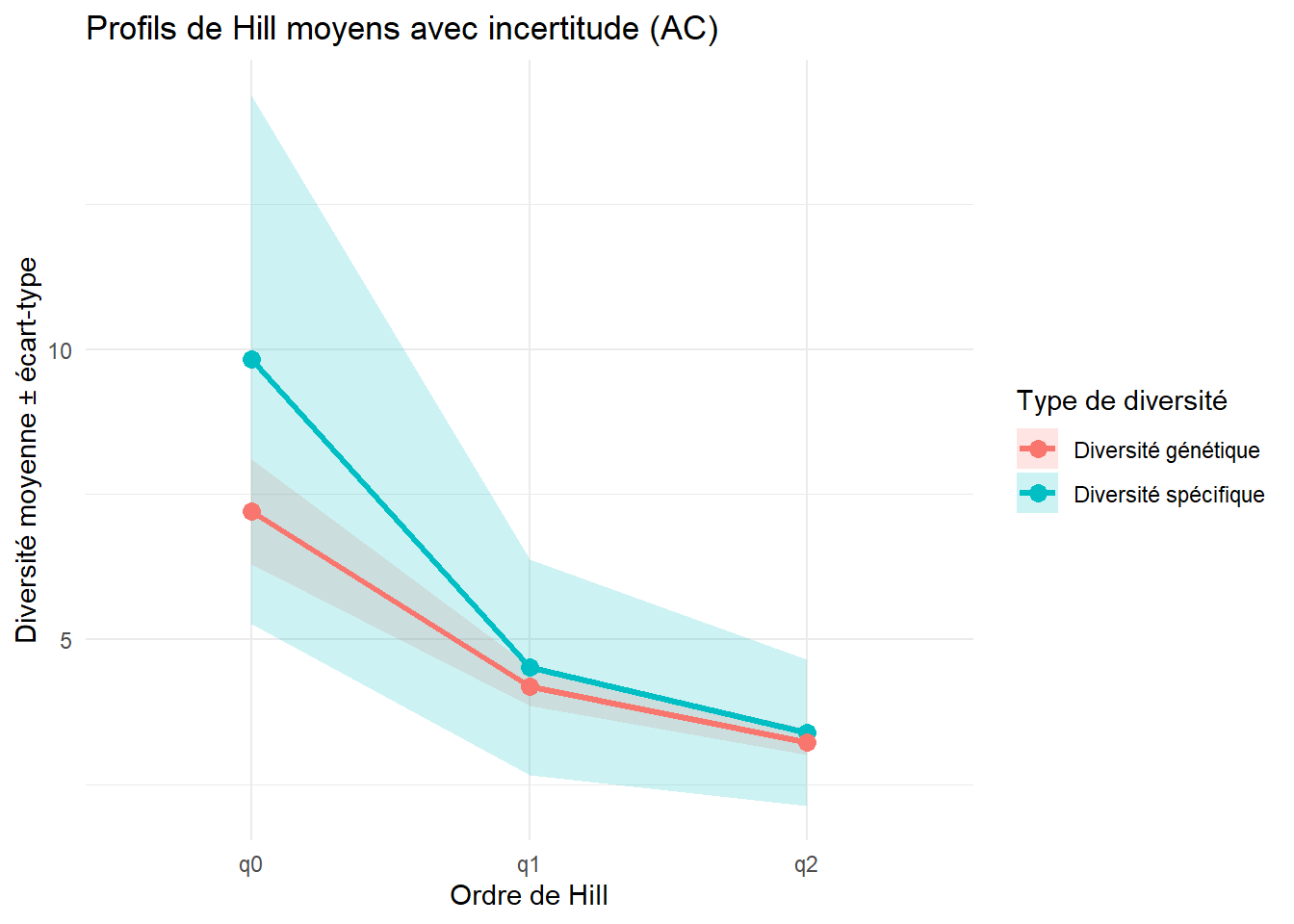
$q1
Pearson's product-moment correlation
data: Hill_gen_vs_spe$q1_sp and Hill_gen_vs_spe$q1_gen
t = 1.6922, df = 22, p-value = 0.1047
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
-0.07419169 0.65332051
sample estimates:
cor
0.3393617
$q2
Pearson's product-moment correlation
data: Hill_gen_vs_spe$q2_sp and Hill_gen_vs_spe$q2_gen
t = 1.2491, df = 22, p-value = 0.2248
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
-0.1629770 0.5985951
sample estimates:
cor
0.2573382 `geom_smooth()` using formula = 'y ~ x'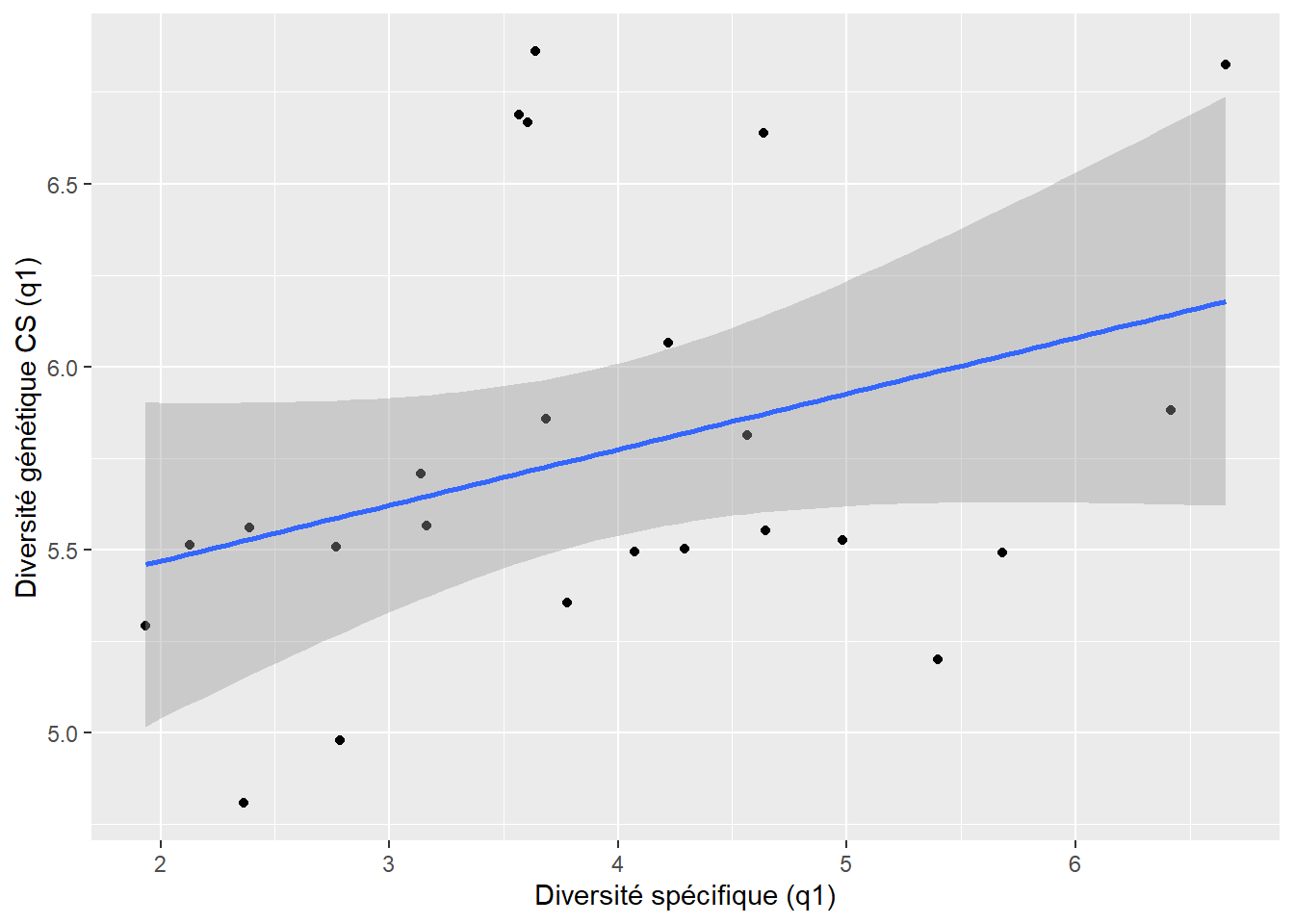
$q1
Pearson's product-moment correlation
data: Hill_gen_vs_spe$q1_sp and Hill_gen_vs_spe$q1_gen
t = 0.24284, df = 27, p-value = 0.81
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
-0.3253889 0.4062391
sample estimates:
cor
0.04668417
$q2
Pearson's product-moment correlation
data: Hill_gen_vs_spe$q2_sp and Hill_gen_vs_spe$q2_gen
t = -0.27012, df = 27, p-value = 0.7891
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
-0.4106074 0.3206932
sample estimates:
cor
-0.05191427 `geom_smooth()` using formula = 'y ~ x'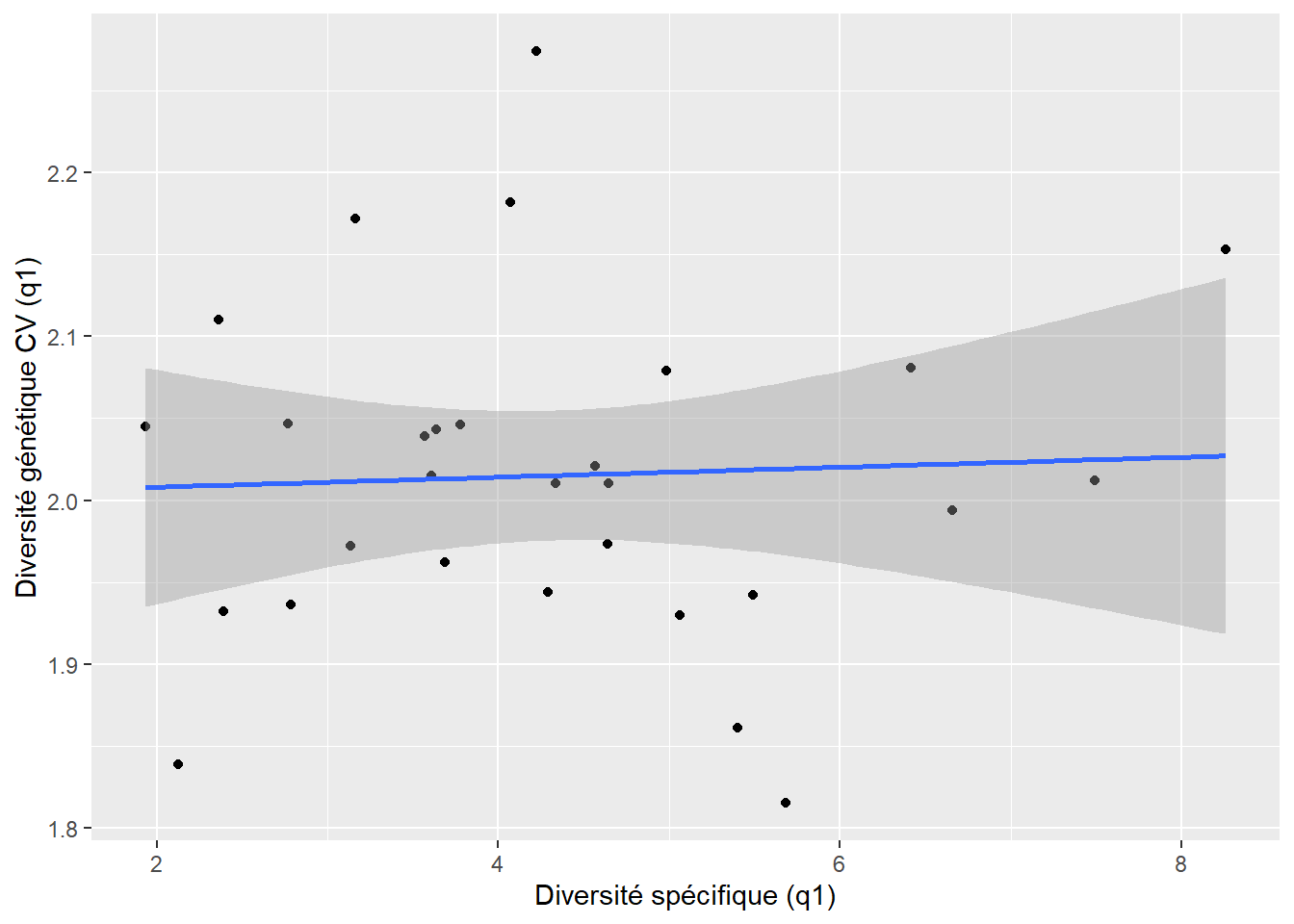
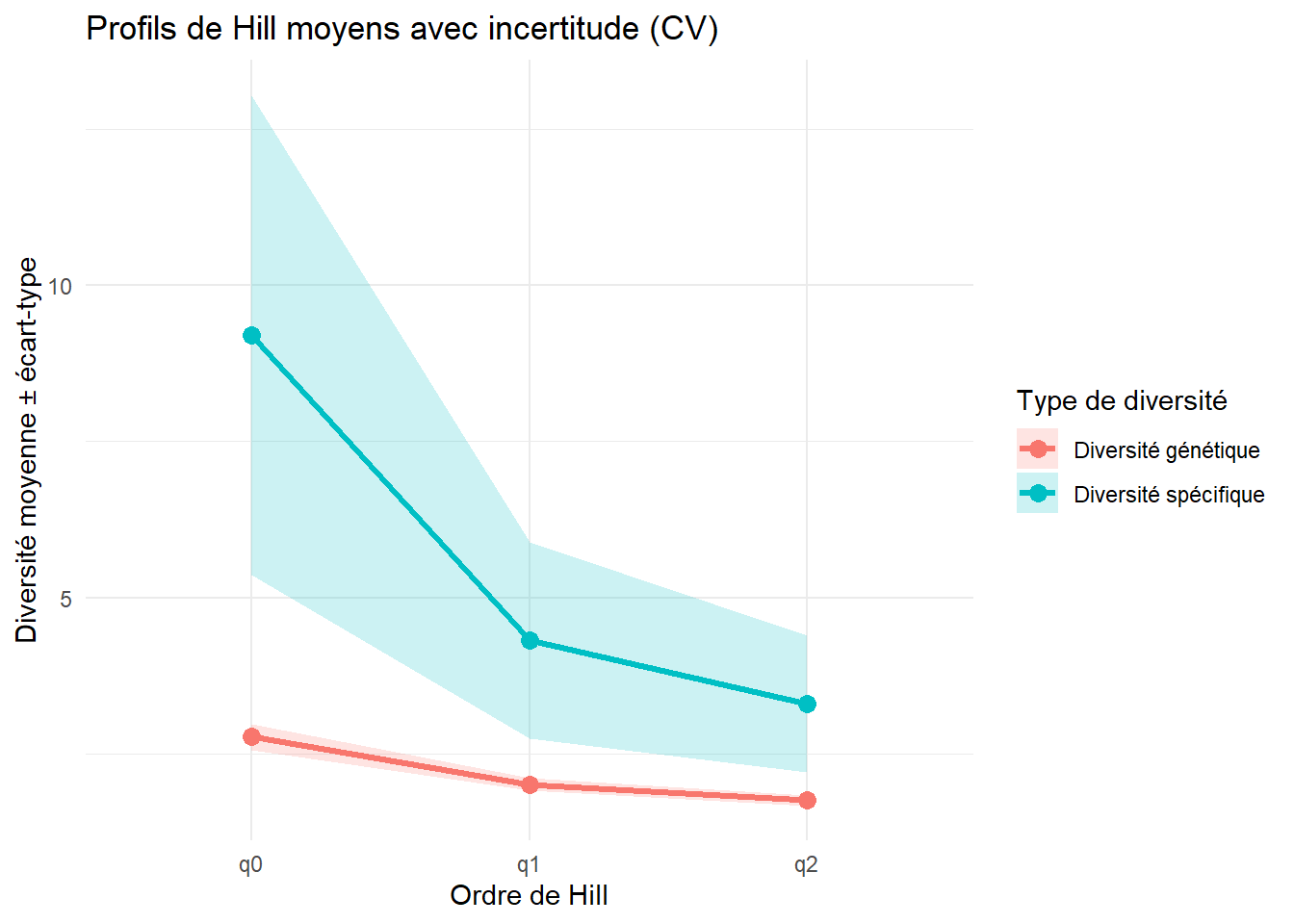
$q1
Pearson's product-moment correlation
data: Hill_gen_vs_spe$q1_sp and Hill_gen_vs_spe$q1_gen
t = 0.36323, df = 26, p-value = 0.7194
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
-0.3102459 0.4326628
sample estimates:
cor
0.07105539
$q2
Pearson's product-moment correlation
data: Hill_gen_vs_spe$q2_sp and Hill_gen_vs_spe$q2_gen
t = 0.17312, df = 26, p-value = 0.8639
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
-0.3434936 0.4019206
sample estimates:
cor
0.03393167 `geom_smooth()` using formula = 'y ~ x'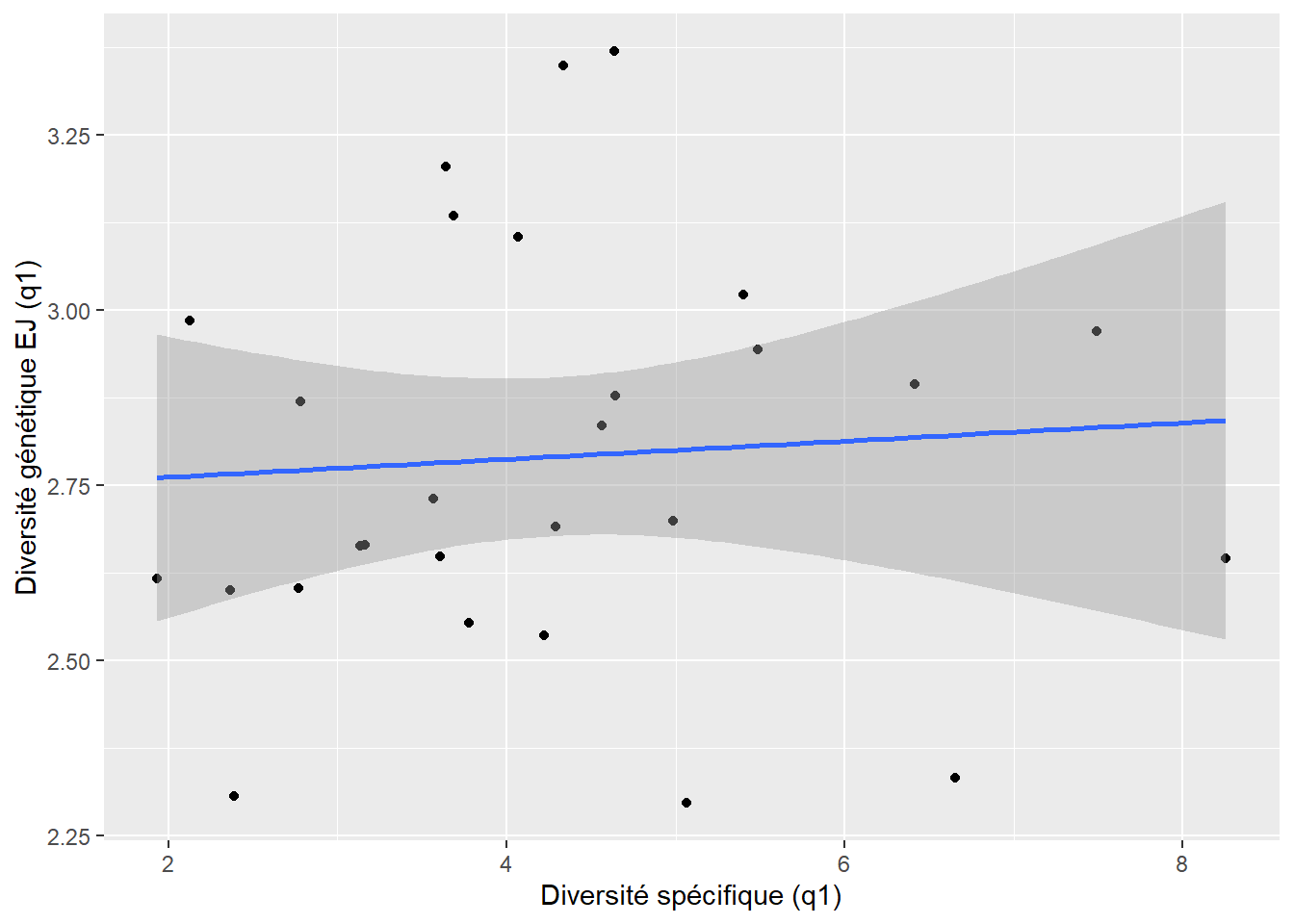
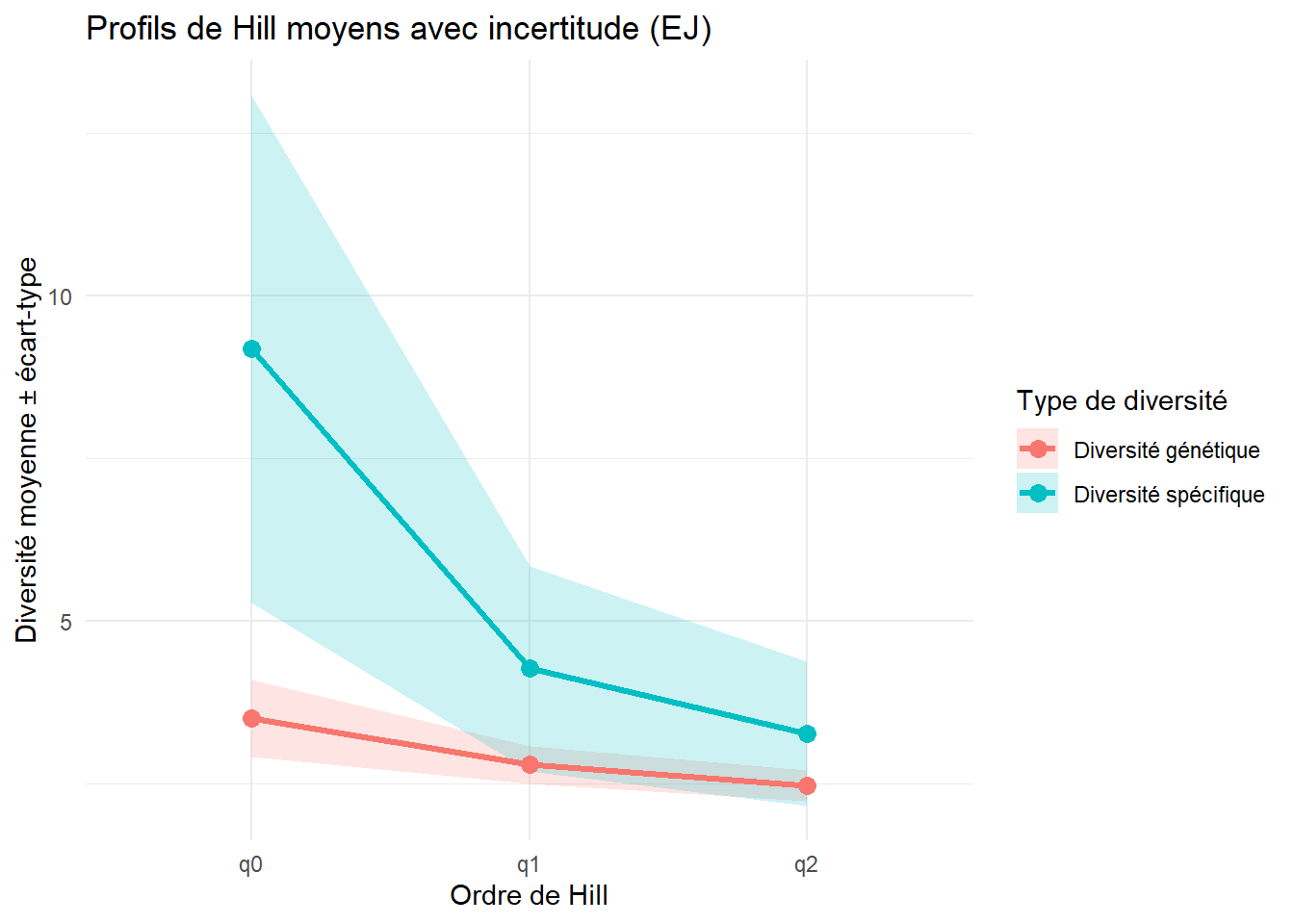
$q1
Pearson's product-moment correlation
data: Hill_gen_vs_spe$q1_sp and Hill_gen_vs_spe$q1_gen
t = 0.58221, df = 27, p-value = 0.5653
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
-0.2660116 0.4591190
sample estimates:
cor
0.1113501
$q2
Pearson's product-moment correlation
data: Hill_gen_vs_spe$q2_sp and Hill_gen_vs_spe$q2_gen
t = 0.26128, df = 27, p-value = 0.7959
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
-0.3222168 0.4091937
sample estimates:
cor
0.05021955 `geom_smooth()` using formula = 'y ~ x'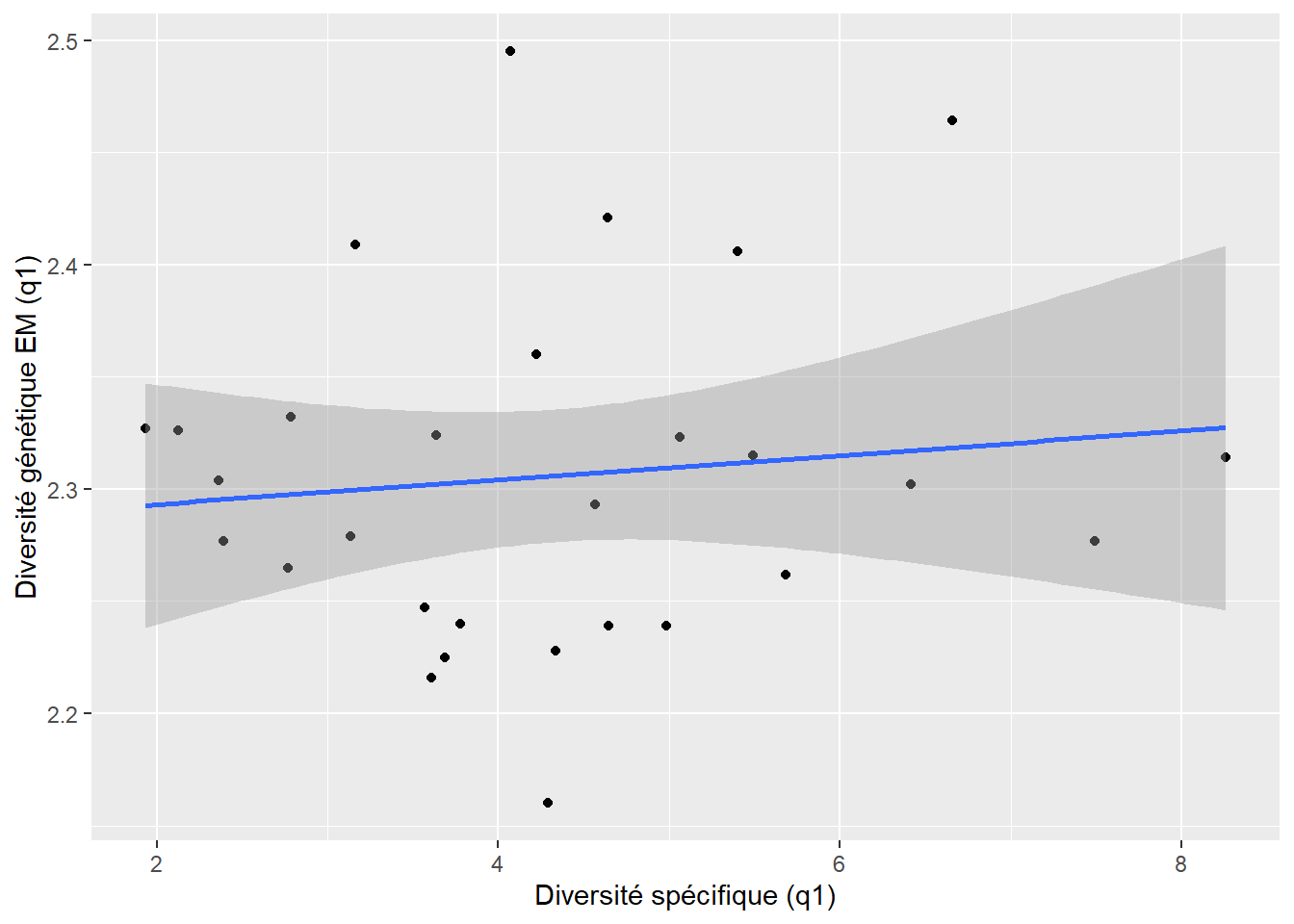
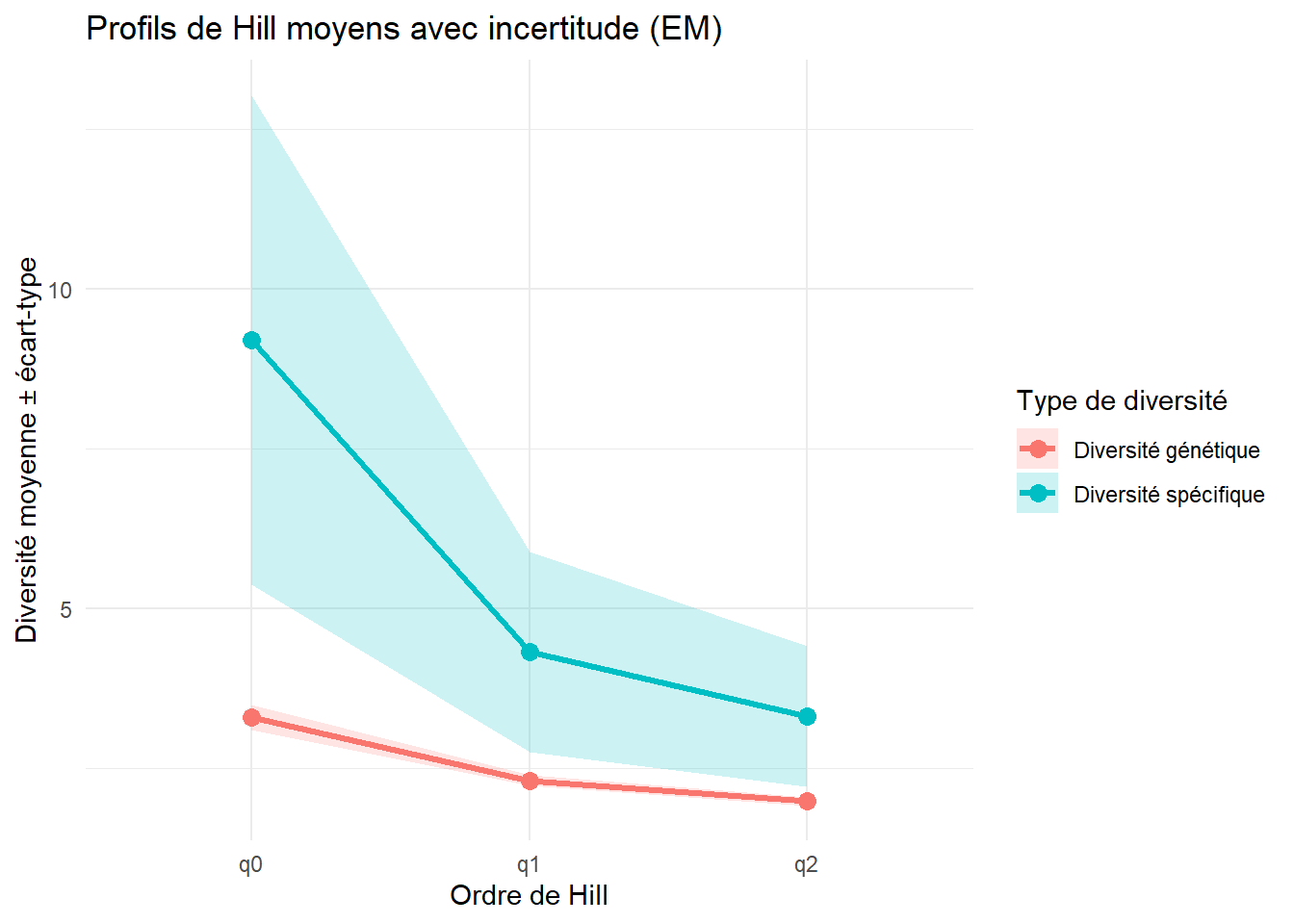
$q1
Pearson's product-moment correlation
data: Hill_gen_vs_spe$q1_sp and Hill_gen_vs_spe$q1_gen
t = -1.6546, df = 27, p-value = 0.1096
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
-0.60287601 0.07098839
sample estimates:
cor
-0.3034113
$q2
Pearson's product-moment correlation
data: Hill_gen_vs_spe$q2_sp and Hill_gen_vs_spe$q2_gen
t = -1.8419, df = 27, p-value = 0.0765
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
-0.62418219 0.03691673
sample estimates:
cor
-0.3341095 `geom_smooth()` using formula = 'y ~ x'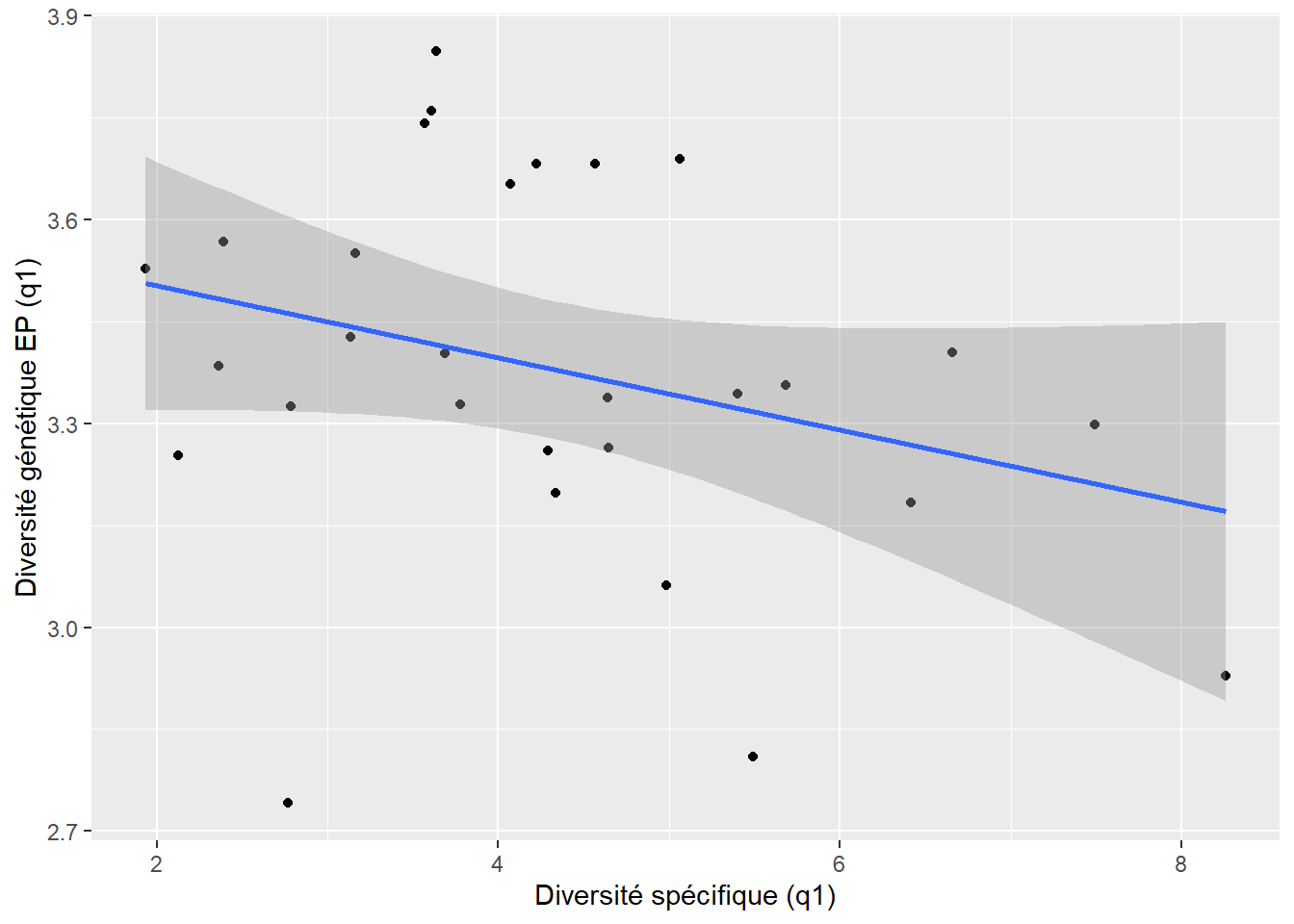
$q1
Pearson's product-moment correlation
data: Hill_gen_vs_spe$q1_sp and Hill_gen_vs_spe$q1_gen
t = 3.1775, df = 26, p-value = 0.003809
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
0.1941013 0.7533173
sample estimates:
cor
0.5288795
$q2
Pearson's product-moment correlation
data: Hill_gen_vs_spe$q2_sp and Hill_gen_vs_spe$q2_gen
t = 2.5528, df = 26, p-value = 0.0169
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
0.08955036 0.70328779
sample estimates:
cor
0.447671 `geom_smooth()` using formula = 'y ~ x'